Specific Aims
Dyslexic people have difficulties in reading, specifically characterized by trouble reading and deciphering language. During reading, sensory hubs in the brain serve as a centralized traffic controller systems to gather different sensory information. For a Dyslexic person, these hubs are poorly connected. [1] Dyslexics also have greater abilities for problem solving possibly due to their enhanced creativity. [2] Interestingly, creativity is also related to neuronal connectivity of the hubs in the brain. One gene associated with dyslexia is DYX1C1 which is a dynein axonemal assembly factor [3]. DYX1C1 is essential for primary cilia construction necessary for neuronal signaling [4]. GAP: Behavioral studies have showed that there is a strong correlation between dyslexia and creativity, yet the molecular and physiological explanation wasn't identified.
|
Goal: My goal is to determine the role of DYX1C1 in creativity as it pertains to cilia signals in the brain. I hypothesize that loss of DYX1C1 leads to differences in the activity of primary cilia essential for creativity. I will use Mus musculus (Mouse) as a model organism, because DYX1C1 knockouts comparing to wild type have significant different brain sizes. Learning, memory and behavior can also be easily determined in both wild type and mutant mice [5]. Mouse can also be easily assayed using a brain fMRI scan, and constructing brain connectivity diagrams. Answering this question helps us realize potentials and talents of dyslexics better.
|
Aim 1: Identify conserved amino acids of DYX1C1 necessary for creativity.
|
Rationale:Different DYX1C1 mutation may cause various defects in primary cilia signaling, which can lead to different forms of dyslexia correlating to creativity. Approach: I will BLAST for homologs and align with Clustal Omega to identify DYX1C1 highly conserved amino acids. I will use CRISPR to edit conserved regions, and confirm different loss of connectivity in the brain with fMRI scan and brain connectivity construct. I will then confirm dyslexia and creativity in the mutated mouse separately in a learning maze and creativity assay. I hypothesize that mice with conserved amino acid knockouts that displayed typical dyslexic brain connectivity loss will be the higher creativity mice. |
Aim 2: Identify genes that are differentially transcribed in DYX1C1 deficient mice with different levels of creativity.
|
Rationale: I want to identify gene ontology groups associated with the mice with low and high levels of creativity in the DYX1C1 mutants. Approach: I will use RNA-Seq on mutated DYX1C1 and wild type mice. I will then use GO to sort the genes related to cilia function, brain function and brain connectivity. I will then knock out the genes that have both three or most three GO groups. I will perform a fMRI scan for creativity assay. I hypothesize that gene involved in primary cilia function will also show high levels of creativity. |
Aim 3: Identify novel proteins that are important creativity in dyslexic mice
|
Rationale: Goal is to identify novel cilia proteins important for creativity.
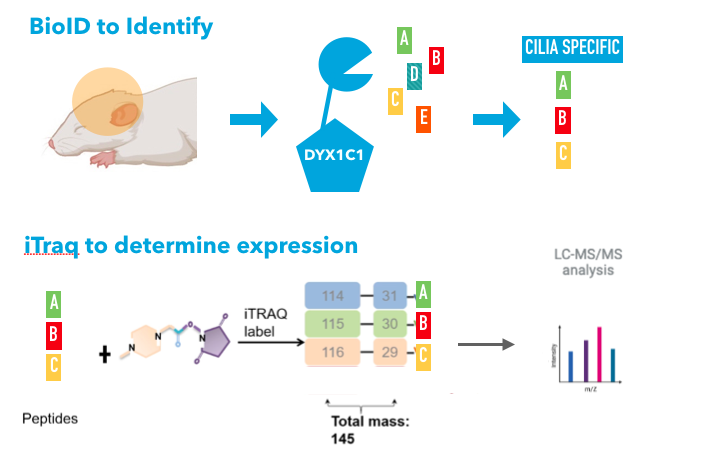
Approach: I will use BioID to tag DYX1C1 and select those that are cilia specific interacting proteins. I will then use iTRAQ to determine which of these proteins are highly expressed between DYX1C1 mutant and wild type. I will also verify creativity levels by performing fMRI scans on BioID proteins that when mutated results in high levels of creativity. I hypothesize that one of these proteins will function in the primary cilia |
Specific aims draft 1: read me!
Specific aims draft 2: read me!
Specific aims draft 3: read me!
Specific aims final: read me!
Specific aims draft 2: read me!
Specific aims draft 3: read me!
Specific aims final: read me!
Citation
[1]Learning difficulties due to poor connectivity, not specific brain regions, study shows. (2020, February 27). Retrieved from https://www.cam.ac.uk/research/news/learning-difficulties-due-to-poor-connectivity-not-specific-brain-regions-study-shows
[2]Cancer, A., & Catholic University of the Sacred Heart of Milan. (n.d.). The alleged link between creativity and dyslexia: Identifying the specific process in which dyslexic students excel. Retrieved from https://www.tandfonline.com/doi/full10.1080/23311908.2016.1190309
[3] Tarkar, A., Loges, N. T., Slagle, C. E., Francis, R., Dougherty, G. W., Tamayo, J. V., Shook, B., Cantino, M., Schwartz, D., Jahnke, C., Olbrich, H., Werner, C., Raidt, J., Pennekamp, P., Abouhamed, M., Hjeij, R., Köhler, G., Griese, M., Li, Y., Lemke, K., … Omran, H. (2013). DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nature genetics, 45(9), 995–1003.https://doi.org/10.1038/ng.2707
[4] Munzer, T., Hussain, K., & Soares, N. (2020). Dyslexia: neurobiology, clinical features, evaluation and management. Translational pediatrics, 9(Suppl 1), S36–S45. https://doi.org/10.21037/tp.2019.09.07
[5]Rendall, A. R., Tarkar, A., Contreras-Mora, H. M., LoTurco, J. J., & Fitch, R. H. (2017, September). Deficits in learning and memory in mice with a mutation of the candidate dyslexia susceptibility gene Dyx1c1. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4646737/
[1]Learning difficulties due to poor connectivity, not specific brain regions, study shows. (2020, February 27). Retrieved from https://www.cam.ac.uk/research/news/learning-difficulties-due-to-poor-connectivity-not-specific-brain-regions-study-shows
[2]Cancer, A., & Catholic University of the Sacred Heart of Milan. (n.d.). The alleged link between creativity and dyslexia: Identifying the specific process in which dyslexic students excel. Retrieved from https://www.tandfonline.com/doi/full10.1080/23311908.2016.1190309
[3] Tarkar, A., Loges, N. T., Slagle, C. E., Francis, R., Dougherty, G. W., Tamayo, J. V., Shook, B., Cantino, M., Schwartz, D., Jahnke, C., Olbrich, H., Werner, C., Raidt, J., Pennekamp, P., Abouhamed, M., Hjeij, R., Köhler, G., Griese, M., Li, Y., Lemke, K., … Omran, H. (2013). DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nature genetics, 45(9), 995–1003.https://doi.org/10.1038/ng.2707
[4] Munzer, T., Hussain, K., & Soares, N. (2020). Dyslexia: neurobiology, clinical features, evaluation and management. Translational pediatrics, 9(Suppl 1), S36–S45. https://doi.org/10.21037/tp.2019.09.07
[5]Rendall, A. R., Tarkar, A., Contreras-Mora, H. M., LoTurco, J. J., & Fitch, R. H. (2017, September). Deficits in learning and memory in mice with a mutation of the candidate dyslexia susceptibility gene Dyx1c1. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4646737/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.