Introduction
Dyslexia is a learning disorder characterized by trouble reading and deciphering language. Dyslexics have a wide range of symptoms including flipping of words, association to the wrong phonology, and unable to identify specific letters.
There has been a lot of misunderstanding in the cause of dyslexia in the past century. People thought it is caused by certain part of the brain, but it is in fact caused by low connectivity of the brain. [1] By measuring connectivities in the brain using fMRI, researchers find that dyslexics differs from normal people by loss of connections from various regions. [2]
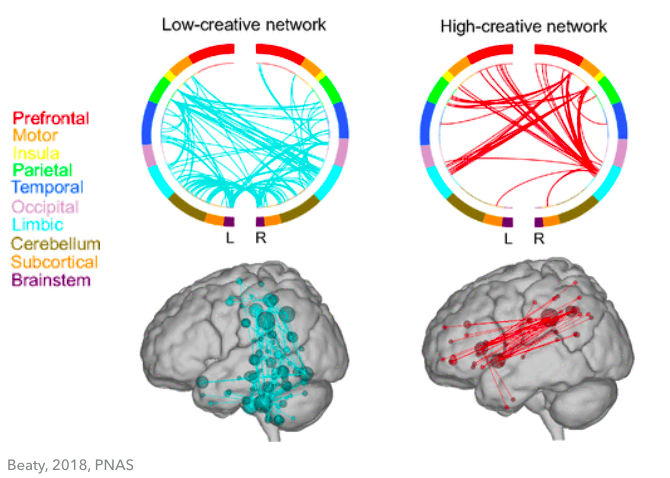
Dyslexics are known to be creative thinkers and problem solvers. Interestingly, another group of researchers found that high creativity people comparing to low creativity people also have less connections in the brain, and high creativity people have connections from the prefrontal lobe. [3]
Is there an explanation physiologically or genetically that dyslexics are more creative?
There has been a lot of misunderstanding in the cause of dyslexia in the past century. People thought it is caused by certain part of the brain, but it is in fact caused by low connectivity of the brain. [1] By measuring connectivities in the brain using fMRI, researchers find that dyslexics differs from normal people by loss of connections from various regions. [2]
Dyslexics are known to be creative thinkers and problem solvers. Interestingly, another group of researchers found that high creativity people comparing to low creativity people also have less connections in the brain, and high creativity people have connections from the prefrontal lobe. [3]
Is there an explanation physiologically or genetically that dyslexics are more creative?
DYX1C1
To understand this relationship, I will use DYX1C1, Dyslexia Susceptibility 1 Candidate Gene 1. DYX1C1 are in the "feet" of neurons which are in charge of moving the cells around as well as transmitting signals. DYX1C1 assembles the outer and inner dynein in cilia.[4] Could mutation in DYX1C1 changes signal transduction of cilia? My goal is to determine if the dyslexia gene DYX1C1 plays a role in creativity through cilia signaling in the brain.
Model Organism
I will use mouse as model organism because
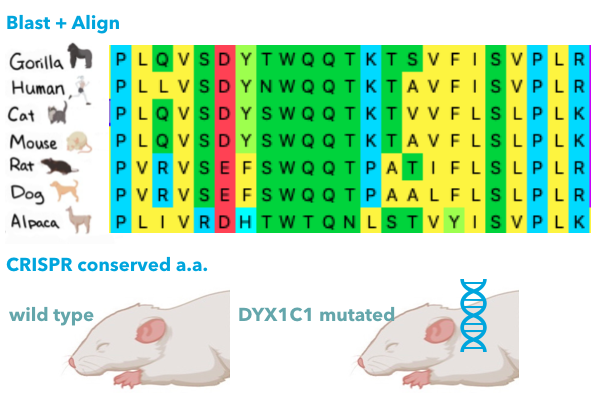
1) DYX1C1 mutated mouse have a significant brain shape difference with wild type
2) There are well established assay for DYX1C1C mutants in memory and learning test. In this study [5], I will use the Y-shaped maze. I will train mouse to recognize the star shape, and mouse will receive a reward if choose the correct path. Dyslexic mouse will have trouble recognizing the right shape.
3) Mouse brain functional structure are similar to human. So it will be easy to construct the brain connectivity map.
1) DYX1C1 mutated mouse have a significant brain shape difference with wild type
2) There are well established assay for DYX1C1C mutants in memory and learning test. In this study [5], I will use the Y-shaped maze. I will train mouse to recognize the star shape, and mouse will receive a reward if choose the correct path. Dyslexic mouse will have trouble recognizing the right shape.
3) Mouse brain functional structure are similar to human. So it will be easy to construct the brain connectivity map.
Specific Aims
Specific aims 1:
I will first try to identify conserved amino acids in the DYX1C1 sequence necessary for creativity, because mutation on specific sites on DYX1C1 affects dyslexia differently, and I wonder if it affects creativity differently. I will BLAST for human DYX1C1 homologs and align the sequences by Clustal Omega. I will use CRISPR to edit sites that are well conserved. I will test if the created mice are dyslexic by their head shape and with Y-shaped maze test. I will then measure how creative these mice are. I hypothesize that the mice show most trouble in the Y-shaped maze test will be determined as more creative by the Y-shaped maze test.
Specific aims 2:
Next, I want to identify genes that are differentially transcribed in DYX1C1 deficient mice, and how they correlate to high and low creativity. I will use RNA-Seq on DYX1C1 and wild type mice, and sort the genes relating to cilia function, neuronal connectivity and brain function by Gene Ontology. I will choose the genes that are with all 3 terms and knock out on mice. I will perform a creativity assay and fMRI and I hypothesize that the ones show high creativity have genes involved in primary cilia function knocked out.
Specific aims 3:
Finally I want to look at if there are novel cilia proteins essential for creativity, so my aim is to identify novel proteins important in creativity. I will use BioID to tag DYX1C1 to probe for proteins that come close to DYX1C1. I will select the ones that are cilia specific. I will then use iTRAQ to determine which of these proteins are highly expressed between DYX1C1 mutant and wild type. I will create a mutant of those highly expressed to verify their dyslexia and creativity. I hypothesize that these proteins are closely related to primary cilia.
Conclusion
Both Dyslexia and high creativity are related to loss of brain connectivity. The dyslexia relating gene DYX1C1 specifically constructs the dynein in primary cilia who is in charge of signal transduction. Mutation in the gene DYX1C1 may result in loss of brain connectivity. Here I used DYX1C1 to explore how dyslexia affects creativity through specific amino acids, DYX1C1 transcriptomics and DYX1C1 proteomics .
In the future...
In the future, it would be interesting to look at how other dyslexia causing genes, such as DCDC2 or KIAA0319 related to creativity. It will also be interesting to see how dyslexia in different languages affect how people think creatively.
Moreover, I hope to use this study and website to help people understand dyslexics better, and realize their potential in many things outside of school.
Moreover, I hope to use this study and website to help people understand dyslexics better, and realize their potential in many things outside of school.
[1]Learning difficulties due to poor connectivity, not specific brain regions, study shows. (2020, February 27). Retrieved from https://www.cam.ac.uk/research/news/learning-difficulties-due-to-poor-connectivity-not-specific-brain-regions-study-shows
[2]Cancer, A., & Catholic University of the Sacred Heart of Milan. (n.d.). The alleged link between creativity and dyslexia: Identifying the specific process in which dyslexic students excel. Retrieved from https://www.tandfonline.com/doi/full10.1080/23311908.2016.1190309
[3]Wang, H.-L. S., Wang, N. Y.-H., & Yeh, F.-C. (2019). Specifying the diffusion MRI connectome in Chinese-speaking children with developmental dyslexia and auditory processing deficits. Pediatrics & Neonatology, 60(3), 297–304. doi: 10.1016/j.pedneo.2018.07.016
[4] Tarkar, A., Loges, N. T., Slagle, C. E., Francis, R., Dougherty, G. W., Tamayo, J. V., Shook, B., Cantino, M., Schwartz, D., Jahnke, C., Olbrich, H., Werner, C., Raidt, J., Pennekamp, P., Abouhamed, M., Hjeij, R., Köhler, G., Griese, M., Li, Y., Lemke, K., … Omran, H. (2013). DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nature genetics, 45(9), 995–1003.https://doi.org/10.1038/ng.2707
[5]Rendall, A. R., Tarkar, A., Contreras-Mora, H. M., LoTurco, J. J., & Fitch, R. H. (2017). Deficits in learning and memory in mice with a mutation of the candidate dyslexia susceptibility gene Dyx1c1. Brain and language, 172, 30–38. https://doi.org/10.1016/j.bandl.2015.04.008
[6]Learning and Memory Test. Stanford Medicine. Retreived from https://med.stanford.edu/sbfnl/services/bm/lm.html
[2]Cancer, A., & Catholic University of the Sacred Heart of Milan. (n.d.). The alleged link between creativity and dyslexia: Identifying the specific process in which dyslexic students excel. Retrieved from https://www.tandfonline.com/doi/full10.1080/23311908.2016.1190309
[3]Wang, H.-L. S., Wang, N. Y.-H., & Yeh, F.-C. (2019). Specifying the diffusion MRI connectome in Chinese-speaking children with developmental dyslexia and auditory processing deficits. Pediatrics & Neonatology, 60(3), 297–304. doi: 10.1016/j.pedneo.2018.07.016
[4] Tarkar, A., Loges, N. T., Slagle, C. E., Francis, R., Dougherty, G. W., Tamayo, J. V., Shook, B., Cantino, M., Schwartz, D., Jahnke, C., Olbrich, H., Werner, C., Raidt, J., Pennekamp, P., Abouhamed, M., Hjeij, R., Köhler, G., Griese, M., Li, Y., Lemke, K., … Omran, H. (2013). DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nature genetics, 45(9), 995–1003.https://doi.org/10.1038/ng.2707
[5]Rendall, A. R., Tarkar, A., Contreras-Mora, H. M., LoTurco, J. J., & Fitch, R. H. (2017). Deficits in learning and memory in mice with a mutation of the candidate dyslexia susceptibility gene Dyx1c1. Brain and language, 172, 30–38. https://doi.org/10.1016/j.bandl.2015.04.008
[6]Learning and Memory Test. Stanford Medicine. Retreived from https://med.stanford.edu/sbfnl/services/bm/lm.html
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.